Bioinformatics on the Raspberry Pi 4
Background
The Raspberry Pi is a widely used single board computer (SBC). This is a small device with most of the components for a full computer built into the board including the CPU, memory, ports and hardware controllers. They are designed to be cheap and portable. These are used to all sorts of applications like home web servers, robotic controllers and so on. Until recently these were not very useful as desktop computers but with the release of the latest version, the Pi 4, it is now feasible to use this device as an actual computer. This is mainly because you can now buy a Pi with 4GB of memory along with a faster CPU. You also need to buy an SD card and power supply. The 4GB version is £55. Also the Pi doesn’t run the desktop version of Windows so be prepared to use Linux.
Benchmark
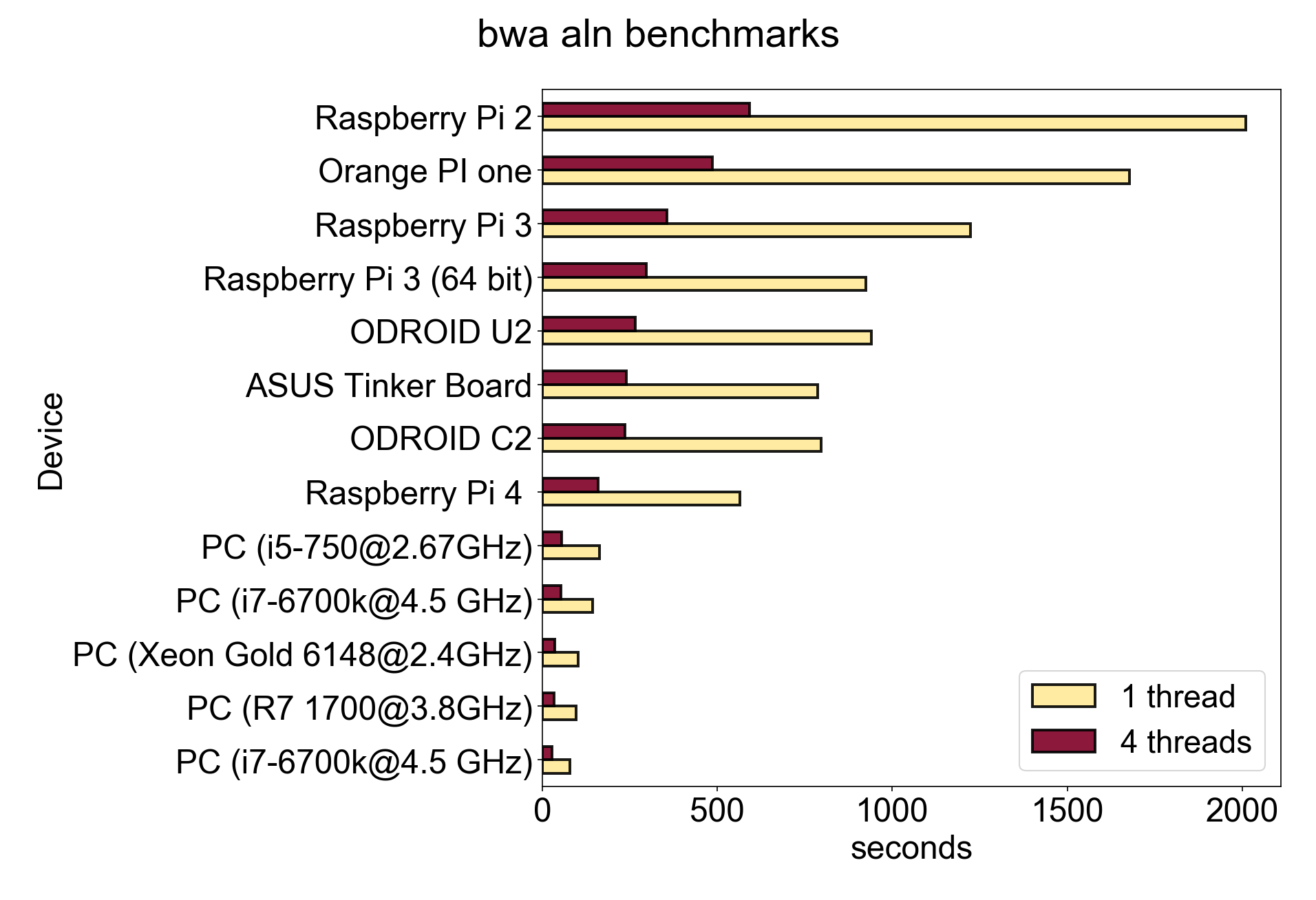
So how does the Pi 4 stack up when compared to other computers when performing a typical bioinformatic task? It turns out that short read alignment is good for doing a so called ‘non-synthetic’ benchmark, as meaningful work is done, and IO, Memory and CPU is all used. In other words it’s a realistic test. Here we show how bwa aln (a tool for mapping reads against a genome) compares between the Pi and other SBCs and PCs. These measurements (except the Pi 4 which I tested myself) are taken from site that did these tests themselves, linked below. The test aligns some short reads to an ecoli genome using either 1 or 4 threads. The bar chart below shows the times taken for each computer (shorter is better). The new Pi is superior to the other SBCs shown. It cannot compare with a full PC grade CPU like an Intel i5 or a Ryzen processor of course. Also those processors now have 12 or more threads so would scale much higher for multi-threaded use. However for many applications this performance would be good enough. There are other SBCs that would beat it too but they are more expensive.
Availibility of software
Linux software is largely installed from repositories. So Ubuntu has repositories with all their compiled software. Raspbian is the operating system made for the Pi. It’s repositories are based on Debian and are fairly comprehensive. Though some software won’t be available if it requires a 64 bit system as Raspian is 32 bit by default. In short most programs will be available on the Pi and you can even download the source and compile them if required. This will likely be the case for more specialised scientific tools. Standard tools work fine on the Pi including the Python and R stacks. If you ever used Jupyter notebooks or R studio they work fine on the Pi.
Cool it!
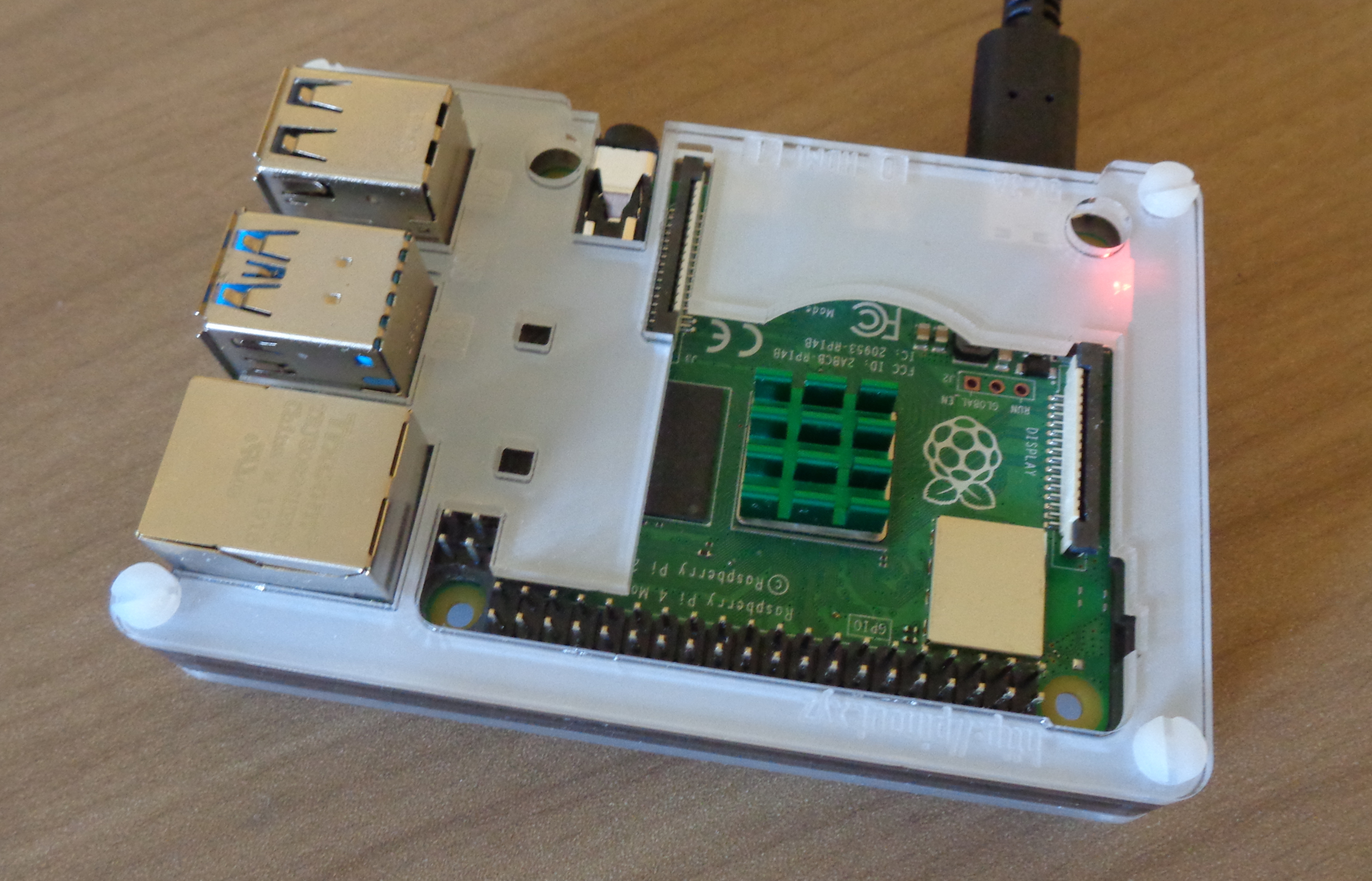
Heat is now an issue with the Pi 4 compared to the older models and it will ‘throttle’ at high temperatures. This just means the CPU has to slow itself down to prevent overheating and be slower to run some tasks unless you can avoid this happening. You can add a fan but the minimum you should do is get a small heat sink and a case that is open at the top allowing some air flow. Something like the one shown in the image above.
Using a Pi for your work
It is in fact feasible to use the Pi 4 for your everyday work, including bioinformatic workflows. Though obviously you must deal with the limitations of speed and memory. It can multitask so you can run programs and still use the desktop. It may not replace a good laptop but remember that you can also use the pi ‘headless’, that is without a display. You then login via ssh in a terminal and run programs. In this way you could use a Pi as an extra box doing some computations while you free up your laptop for other work. Pis are used like this in computer clusters where many are linked together to perform highly parallel tasks.